- Procrustes Analysis Software For Mac Windows 10
- Procrustes Analysis Software For Mac Windows 10
- Software For Mac Free
- Procrustes Analysis Software For Mac Free
Procrustes Analysis
Compare Landmark Data
Journal of Biomechanics 40 (2007) 437–444 Quantifying biomechanical motion using Procrustes motion analysis Dean C. Adamsa,b, Melinda M. Cerneyc,1 aDepartment of Ecology, Evolution and Organismal Biology, Iowa State University, Ames, IA, USA bDepartment of Statistics, Iowa State University, Ames, IA, USA cHuman Computer Interaction Program, Iowa State University. Based on generalized Procrustes analysis, this function determines a linear transformation (rotation/reflection and scalling) of the points in matrix x to align them to their reference points in matrix xbar. The alignemnt is carried out by minimizing the distance between the points in x and xbar. Thus, generalized Procrustes analysis is a three-mode method of analysis. For some information on algorithms, see ten Berge (1977) To help interpretation, each dimension of the analysis may be summarized in an analysis of variance, partitioning the total into terms for the group average and for departures from the group average. D = procrustes(X,Y) determines a linear transformation (translation, reflection, orthogonal rotation, and scaling) of the points in matrix Y to best conform them to the points in matrix X. The goodness-of-fit criterion is the sum of squared errors. Procrustes returns the minimized value of this dissimilarity measure in d. Procrustes analysis takes a set of landmark points, encoded by geometric coordinates (either in 2D or in 3D), and computes a transformation to align them to a reference face 9.
The
procrustes function analyzes the distribution of a set of shapes using Procrustes analysis. This analysis method matches landmark data (geometric locations representing significant features in a given shape) to calculate the best shape-preserving Euclidean transformations. These transformations minimize the differences in location between compared landmark data. Procrustes analysis is also useful in conjunction with multidimensional scaling. In Construct a Map Using Multidimensional Scaling there is an observation that the orientation of the reconstructed points is arbitrary. Two different applications of multidimensional scaling could produce reconstructed points that are very similar in principle, but that look different because they have different orientations. The
procrustes function transforms one set of points to make them more comparable to the other.Data Input
The
procrustes function takes two matrices as input: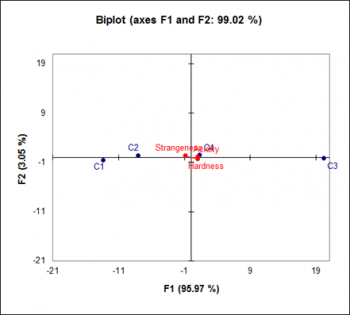
- The target shape matrix X has dimension
n×p, wherenis the number of landmarks in the shape andpis the number of measurements per landmark. - The comparison shape matrix Y has dimension
n×qwithq≤p. If there are fewer measurements per landmark for the comparison shape than the target shape (q<p), the function adds columns of zeros to Y, yielding ann×pmatrix.
The equation to obtain the transformed shape, Z, is
| (1) |
where:
- b is a scaling factor that stretches (b > 1) or shrinks (b < 1) the points.
- T is the orthogonal rotation and reflection matrix.
- c is a matrix with constant values in each column, used to shift the points.
The
procrustes function chooses b, T, and c to minimize the distance between the target shape X and the transformed shape Z as measured by the least squares criterion:Preprocess Data for Accurate Results
Procrustes analysis is appropriate when all
p measurement dimensions have similar scales. The analysis would be inaccurate, for example, if the columns of Z had different scales:- The first column is measured in milliliters ranging from 2,000 to 6,000.
- The second column is measured in degrees Celsius ranging from 10 to 25.
- The third column is measured in kilograms ranging from 50 to 230.
In such cases, standardize your variables by:
- Subtracting the sample mean from each variable.
- Dividing each resultant variable by its sample standard deviation.
Use the
zscore function to perform this standardization.See Also
Related Topics
Geometric Morphometric Analyses of 2D/3D Landmark Data
Read, manipulate, and digitize landmark data, generate shapevariables via Procrustes analysis for points, curves and surfaces, performshape analyses, and provide graphical depictions of shapes and patterns ofshape variation.
Readme
Geomorph is a software package for performing all stages of geometric morphometric shape analysis of 2- and 3-dimensional landmark points, as well as semilandmarks on curves and surfaces, in the R statistical computing environment. This repository is dedicated to providing beta versions between CRAN uploads.
To install the current geomorph R-package from CRAN:
Within R:
install.packages('geomorph') For Mac users: please also install XQuartz from https://www.xquartz.org/. This allows the library(rgl) to function.
To install the current version of geomorph R-package from Github using devtools:
Within R:
install.packages('devtools') devtools::install_github('geomorphR/geomorph', ref = 'Stable', build_vignettes = TRUE)This installs a stable release of the current version of geomorph on CRAN, allowing us to quickly fix errors that slip thorough the cracks and are uploaded with the CRAN version.
To install the Development version (beta) of geomorph R-package from Github using devtools:
Procrustes Analysis Software For Mac Windows 10
Within R:
install.packages('devtools') devtools::install_github('geomorphR/geomorph', ref = 'Develop', build_vignettes = TRUE)NOTE FOR THE PRE-RELEASE (BETA) VERSION: We strongly discourage you from publishing results with this version, unless you check with the authors first.
Functions in geomorph
| Name | Description | |
| buildtemplate | Build 3D surface template | |
| bilat.symmetry | Analysis of bilateral symmetry | |
| compare.multi.evol.rates | Comparing net rates of shape evolution among traits on phylogenies | |
| compare.evol.rates | Comparing net rates of shape evolution on phylogenies | |
| coords.subset | Subset landmark coordinates via a factor | |
| compare.pls | Comparisons of Effect Sizes from Partial Least Squares | |
| arrayspecs | Convert landmark data matrix into array (p x k x n) | |
| combine.subsets | Combine separate landmark configurations | |
| compare.CR | Comparisons of Effect Sizes from Modularity Analyses | |
| define.links | Define links between landmarks | |
| digit.fixed | Digitize 3D landmarks on mesh3d object | |
| digit.curves | Calculate semilandmarks along a curve | |
| digitize2d | Digitize 2D landmarks on .jpg files | |
| digitsurface | Digitize 3D fixed landmarks and surface semilandmarks | |
| geomorph-package | Geometric morphometric analyses for 2D/3D data | |
| globalIntegration | Quantify global integration relative to self-similarity | |
| fixed.angle | Rotate a subset of 2D landmarks to common articulation angle | |
| findMeanSpec | Identify specimen closest to the mean of a set of Procrustes shape variables | |
| gpagen | Generalized Procrustes analysis of points, curves, and surfaces | |
| estimate.missing | Estimate locations of missing landmarks | |
| interlmkdist | Calculate linear distances between landmarks | |
| gm.prcomp | Principal and phylogenetically-aligned components analysis of shape data | |
| editTemplate | Edit 3D template | |
| mshape | Estimate mean shape for a set of aligned specimens | |
| lizards | Dorsal head shape data of lizards | |
| modularity.test | Evaluate the degree of modular signal in shape data | |
| geomorph.data.frame | Create a data frame with shape data | |
| phylo.integration | Quantify phylogenetic morphological integration between two or more sets of variables under Brownian motion | |
| hummingbirds | Landmark data from hummingbird bills (includes sliding semilandmarks on curves) | |
| morphol.disparity | Morphological disparity for one or more groups of specimens | |
| plot.bilat.symmetry | Plot Function for geomorph | |
| plot.evolrate | Plot Function for geomorph | |
| gridPar | Set up parameters for grids, points, and links in plotRefToTarget | |
| phylo.modularity | Evaluate the degree of phylogenetic modular signal in Procrustes shape variables | |
| larvalMorph | Head and tail shapes of larval salamanders | |
| define.modules | Define modules (landmark partitions) | |
| integration.test | Quantify morphological integration between modules | |
| plotGMPhyloMorphoSpace | Deprecated functions in geomorph | |
| plotOutliers | Find potential outliers | |
| physignal | Assessing phylogenetic signal in Procrustes shape variables | |
| plot.mshape | Plot Function for geomorph | |
| define.sliders | Select points to 'slide' along curves | |
| plot.gm.prcomp | Plot Function for geomorph | |
| picknplot.shape | Pick points in geomorph scatterplots to visualize shape variation | |
| procD.lm | Procrustes ANOVA/regression for Procrustes shape variables | |
| print.pls | Print/Summary Function for geomorph | |
| print.combined.set | Print/Summary Function for geomorph | |
| print.procD.lm | Print/Summary Function for geomorph | |
| print.physignal | Print/Summary Function for geomorph | |
| print.compare.pls | Print/Summary Function for geomorph | |
| plot.physignal | Plot Function for geomorph | |
| mosquito | Landmarks on mosquito wings | |
| plot.CR | Plot Function for geomorph | |
| plethodon | Landmark data from Plethodon salamander heads | |
| plethShapeFood | Head shape and food use data from Plethodon salamanders | |
| plot.CR.phylo | Plot Function for geomorph | |
| plotTangentSpace | Deprecated functions in geomorph | |
| plot.gpagen | Plot Function for geomorph | |
| plotRefToTarget | Plot shape differences between a reference and target specimen | |
| plotAllSpecimens | Plot landmark coordinates for all specimens | |
| scallopPLY | 3D scan of a scallop shell from a .ply file in mesh3d format | |
| plotAllometry | Plotting to assist visualization of shape-size covariation (allometry) | |
| scallops | Landmark data from scallop shells | |
| print.gpagen | Print/Summary Function for geomorph | |
| summary.geomorphShapes | Print/Summary Function for geomorph | |
| summary.evolrate1 | Print/Summary Function for geomorph | |
| summary.gpagen | Print/Summary Function for geomorph | |
| plotspec | Plot 3D specimen, fixed landmarks and surface semilandmarks | |
| print.evolrate1 | Print/Summary Function for geomorph | |
| summary.gm.prcomp | Print/Summary Function for geomorph | |
| print.CR | Print/Summary Function for geomorph | |
| print.evolrate | Print/Summary Function for geomorph | |
| plot.pls | Plot Function for geomorph | |
| plot.procD.lm | Plot Function for geomorph | |
| plethspecies | Average head shape and phylogenetic relationships for several Plethodon salamander species | |
| readland.nts | Read landmark data matrix from nts file | |
| warpRefOutline | Creates a 2D outline warped to the mean shape | |
| warpRefMesh | Creates a mesh3d object warped to the mean shape | |
| print.geomorphShapes | Print/Summary function for geomorph | |
| print.gm.prcomp | Print/Summary function for geomorph | |
| readland.shapes | Read landmark data from a shapes object (StereoMorph) | |
| readland.fcsv | Read landmark data matrix from fcsv file | |
| print.morphol.disparity | Print/Summary Function for geomorph | |
| print.CR.phylo | Print/Summary Function for geomorph | |
| read.ply | Read mesh data (vertices and faces) from ply files | |
| readland.tps | Read landmark data from tps file | |
| procD.pgls | Phylogenetic ANOVA/regression for Procrustes shape variables | |
| summary.compare.pls | Print/Summary Function for geomorph | |
| ratland | Landmark data from dataset rat | |
| summary.pls | Print/Summary Function for geomorph | |
| readmulti.nts | Read landmark data from multiple nts files | |
| writeland.tps | Write landmark data to tps file | |
| summary.evolrate | Print/Summary Function for geomorph | |
| print.bilat.symmetry | Print/Summary Function for geomorph | |
| summary.procD.lm | Print/Summary Function for geomorph | |
| pupfish | Landmarks on pupfish | |
| read.morphologika | Read landmark data from Morphologika file(s) | |
| shape.predictor | Shape prediction from numeric predictors | |
| shapeHulls | Update Plots with Convex Hulls for Groups | |
| readmulti.tps | Read and combine multiple tps files | |
| rotate.coords | Rotate or flip landmark or coordinate configurations | |
| summary.CR | Print/Summary Function for geomorph | |
| summary.CR.phylo | Print/Summary Function for geomorph | |
| two.b.pls | Two-block partial least squares analysis for Procrustes shape variables | |
| two.d.array | Convert (p x k x n) data array into 2D data matrix | |
| summary.bilat.symmetry | Print/Summary Function for geomorph | |
| summary.morphol.disparity | Print/Summary Function for geomorph | |
| summary.physignal | Print/Summary Function for geomorph | |
| summary.combined.set | Print/Summary Function for geomorph | |
| No Results! | ||
Procrustes Analysis Software For Mac Windows 10
Vignettes of geomorph
| Name | ||
| figs/PCAplot2D.png | ||
| figs/example_spec2.png | ||
| figs/example_template.png | ||
| figs/fixed.3D.png | ||
| figs/tree.plot.png | ||
| geomorph.PCA.Rmd | ||
| geomorph.assistance.Rmd | ||
| geomorph.digitize3D.Rmd | ||
| geomorph.functions.Rmd | ||
| No Results! | ||
Last month downloads
Details
Software For Mac Free
| License | GPL (>= 2) |
| URL | https://github.com/geomorphR/geomorph |
| Encoding | UTF-8 |
| LazyData | true |
| RoxygenNote | 7.1.0 |
| VignetteBuilder | knitr |
| NeedsCompilation | no |
| Packaged | 2020-06-12 19:54:19 UTC; dcadams |
| Repository | CRAN |
| Date/Publication | 2020-06-12 23:50:07 UTC |
| imports | ape , graphics , grDevices , jpeg , stats , utils |
| suggests | knitr , rmarkdown , testthat |
| depends | R (>= 3.5.0) , rgl , RRPP |
| Contributors | Michael Collyer, Antigoni Kaliontzopoulou |
Procrustes Analysis Software For Mac Free
Include our badge in your README
[](http://www.rdocumentation.org/packages/geomorph) 